Visualization and analysis (tutorial)
This trail shows you how analysis and visualization goes hand in hand in visone. It introduces you to the most common usage scenario: importing data from one or several files, analyzing the network, visualizing the network together with the computed indicators, exporting data and images for further processing or publication.
This trail assumes that you have basic knowledge about how to operate the visone GUI - as explained in the previous trail.
Introducing an exemplary dataset
The data that we use in this trail has been collected in a long-term reseach project about acculturation networks. More information about the project can be found at [1]. Among others, the personal networks of now more than 1,000 immigrants have been collected within this project. Each of the respondents (called ego) provided answers to four types of questions:
- questions about ego, including country of origin, years of residence, age, gender, skin color, reasons for migrating, health, language skills...
- alters a list of persons known to ego (for most networks the number has been fixed to 45)
- questions about alters including country of origin, country of residence, age, skin color, type of relation to ego, ...
- alter-alter ties (undirected) pairs of alters that know each other (according to the respondent)
In this trail we exemplarily analyze one of these personal networks, obtained from interviewing a migrant from the Dominican Republic to the USA. The dataset here contains none of the variables characterizing ego but only the alter characteristics and the alter-alter ties.
More specifically the ties are encoded in an adjacency matrix file Egonet_ties.csv, the alter characteristics in a file Egonet_attributes.csv and both of these two kinds of information are provided (more comfortable and reliable) in a GraphML file Egonet.graphml.
- alter-alter ties Egonet_ties.csv
- alter characteristics Egonet_attributes.csv
- GraphML file Egonet.graphml
Importing networks from adjacency matrix files
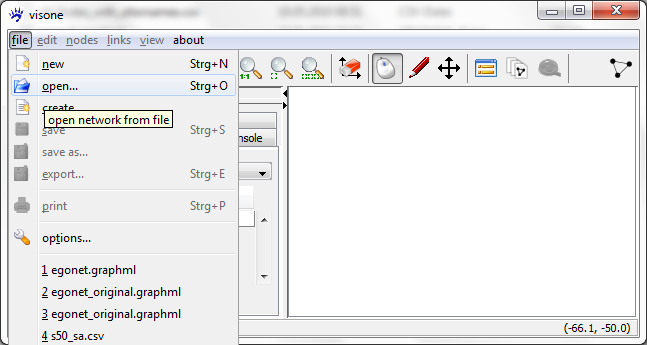
The usual way to get a network into visone is to read it from a local file via the menu file, open
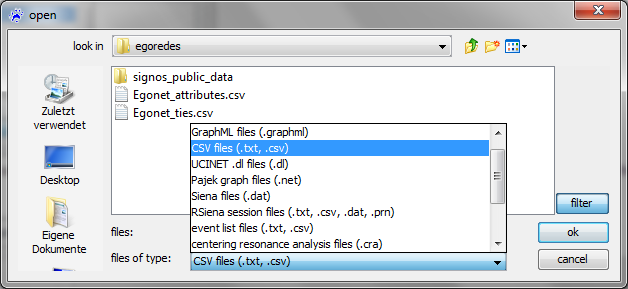
The usual file type to be read by visone is GraphML; GraphML files contain information about nodes and links, about attributes of nodes and links, and about graphical information such as layout, color, or shape. To read GraphML files you select .graphml in the file open dialog (shown below) and click on ok; this is simple, fast, and reliable.
Here, for illustration, we go the hard way and assume that the data are not stored in a GraphML file but in comma-separated-value tables. This very primitive file type can be output from many programs, including statistical software, spread-sheet editors, or other network analysis software. Sometimes you have to deal with this file type.
To open a network from an adjacency matrix file you select the type .txt, .csv in the file open dialog and click on ok. To follow the steps outlined in this trail, select the file egonet_ties.csv.
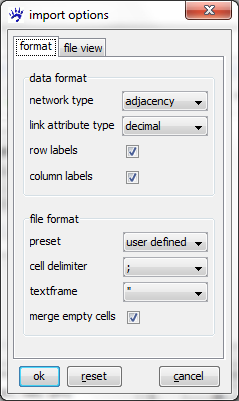
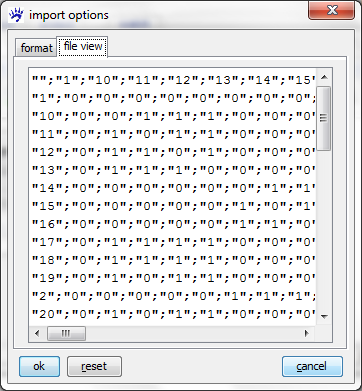
Clicking on ok does not immediatelly open the file. Indeed, in contrast to GraphML, CSV files don't have a self-explaining interpretation; rather the program that has to handle them needs some guidance. Therefore visone opens an import options dialog whose two tabs are shown below.
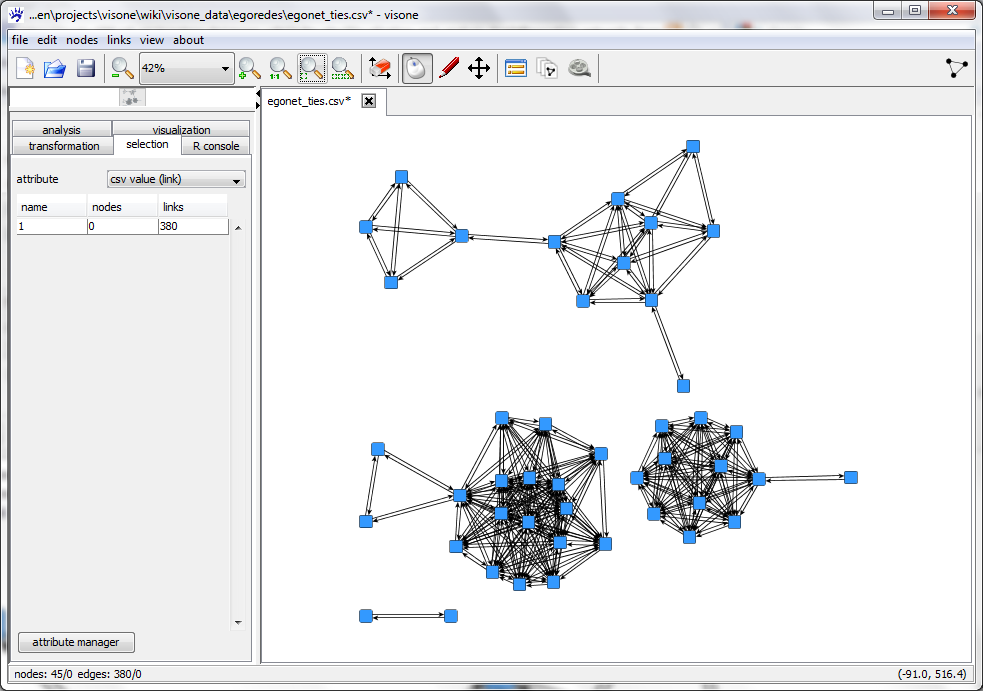
The file view tab shows you (part of) the adjacency matrix encoded in the file to be opened. From this view you can guess, for instance, that different cells in the matrix are delimited by semicolons (;), that row and column labels are present, and some more. For an exhaustive explanation of all options and their meaning see the page on the import options dialog. To continue with this trail, set all options as shown in the format tab above and click on ok. This opens a network looking like this.
The .csv does not contain layout information. The position of the nodes has been determined by the layout algorithm that can be initiated with the quick layout button.
Merging parallel ties
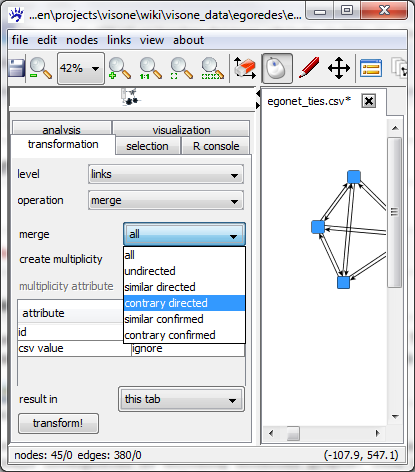
The network above contains for every pair of actors that are connected two anti-parallel ties. This is due to the fact that adjacency matrices are always interpreted as encoding directed graphs. This interpretation is wrong in our example since the tie-generating question was "do actor A and actor B know each other?" which clearly generates an undirected relation. All pairs of anti-parallel directed links can be merged to one undirected link via the transformation tab.
Therefore chose links as the level on which the transformation should be applied, merge as the operation, and chose contrary directed in the drop-down menu right of merge. Clicking on transform! at the bottom of the tab executes the transformation and the network has been transformed into an undirected one with no parallel links.
Since now we have already invested some work in the network, we might save it by clicking on file, save. (The first time we do this we have to assign a name to the network.) Note that the network is saved in GraphML format; indeed only this format guarantees that no information gets lost.