Visualization and analysis (tutorial): Difference between revisions
| Line 70: | Line 70: | ||
[[File:Attribute_manager_import.png]] [[File:Attribute_manager_import_options.png]] | [[File:Attribute_manager_import.png]] [[File:Attribute_manager_import_options.png]] | ||
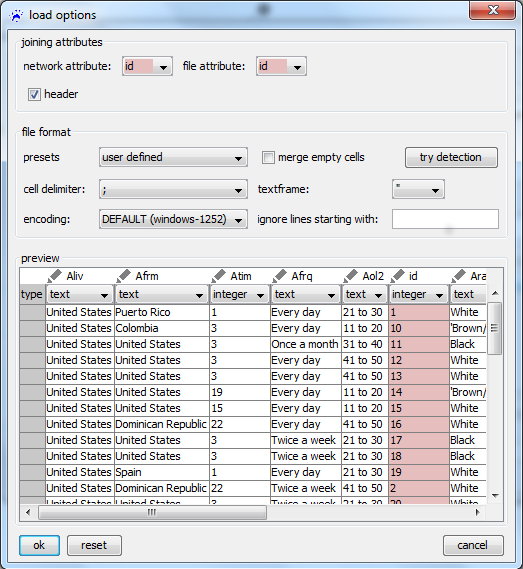
Before clicking on '''apply''' it is very important to correctly set the value in the ''join by'' drop-down menu. This should point to the name of the attribute that identifies the nodes and tells visone which column in the imported CSV file holds these identifiers. Currently you can only select '''id''' but in general there might be several node attributes and the nodes could as well be identified by attributes having a different name than '''id'''. Clicking on '''apply''' opens an import options dialog; setting the options as shown | Before clicking on '''apply''' it is very important to correctly set the value in the ''join by'' drop-down menu. This should point to the name of the attribute that identifies the nodes and tells visone which column in the imported CSV file holds these identifiers. Currently you can only select '''id''' but in general there might be several node attributes and the nodes could as well be identified by attributes having a different name than '''id'''. Clicking on '''apply''' opens an import options dialog; setting the options as shown above - in particular, setting the cell delimiter to semicolon (''';''') - and clicking on '''ok''' imports the attributes. You can see the result by checking the '''values''' radio button on the left hand side of the attribute manager. Then, you should see something similar to the image on the right. | ||
[[File:Attribute_manager_import_result.png]] | [[File:Attribute_manager_import_result.png]] | ||
Revision as of 09:54, 18 February 2011
This trail shows you how analysis and visualization goes hand in hand in visone. It introduces you to the most common usage scenario: importing data from one or several files, analyzing the network, visualizing the network together with the computed indicators, exporting data and images for further processing or publication.
This trail assumes that you have basic knowledge about how to operate the visone GUI - as explained in the previous trail.
Introducing an exemplary dataset
The data that we use in this trail has been collected in a long-term reseach project about acculturation networks. More information about the project can be found at [1]. Among others, the personal networks of now more than 1,000 immigrants have been collected within this project. Each of the respondents (called ego) provided answers to four types of questions:
- questions about ego, including country of origin, years of residence, age, gender, skin color, reasons for migrating, health, language skills...
- alters a list of persons known to ego (for most networks the number has been fixed to 45)
- questions about alters including country of origin, country of residence, age, skin color, type of relation to ego, ...
- alter-alter ties (undirected) pairs of alters that know each other (according to the respondent)
A more detailed description is provided on the page Egoredes (data).
In this trail we exemplarily analyze one of these personal networks obtained from interviewing a migrant from the Dominican Republic to the USA. The dataset here contains none of the variables characterizing ego but only the alter characteristics and the alter-alter ties.
More specifically the ties are encoded in an adjacency matrix file Egonet_ties.csv, the alter characteristics in a file Egonet_attributes.csv. The GraphML file Egonet.graphml contains ties and attributes in a more comfortable and reliable way; the CSV files are provided for the sole purpose of demonstrating how data can be imported from comma-separated value tables.
- alter-alter ties (Egonet_ties.csv)
- alter characteristics (Egonet_attributes.csv)
- GraphML file (Egonet.graphml)
To follow the steps explained in this trail, you should download these three files and save them on your hard disk (right-click and select save link as).
Importing networks from adjacency matrix files
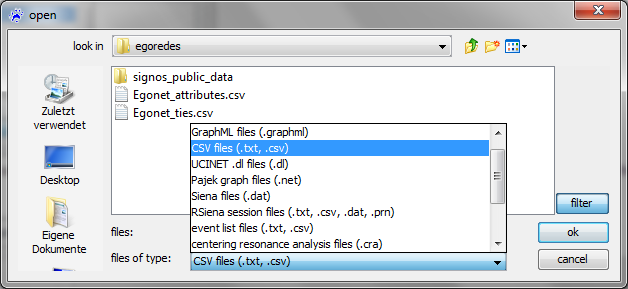
The usual way to get a network into visone is to read it from a local file via the menu file, open
The usual file type to be read by visone is GraphML; GraphML files contain information about nodes and links, about attributes of nodes and links, and about graphical information such as layout, color, or shape. To read GraphML files you select .graphml in the file open dialog (shown below) and click on ok; this is simple, fast, and reliable.
Here, for illustration, we go the hard way and assume that the data are not stored in a GraphML file but in comma-separated-value tables. This very primitive file type can be output from many programs, including statistical software, spread-sheet editors, or other network analysis software. Sometimes you have to deal with this file type.
To open a network from an adjacency matrix file you select the type .txt, .csv in the file open dialog and click on ok. To follow the steps outlined in this trail, select the file egonet_ties.csv.
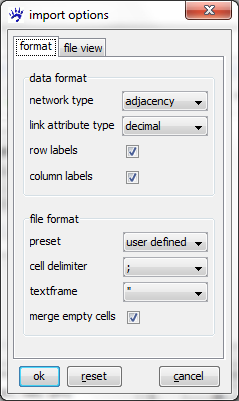
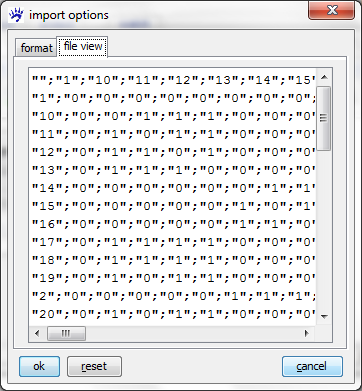
Clicking on ok does not immediatelly open the file. Indeed, in contrast to GraphML, CSV files don't have a self-explaining interpretation; rather the program that has to handle them needs some guidance. Therefore visone opens an import options dialog whose two tabs are shown below.
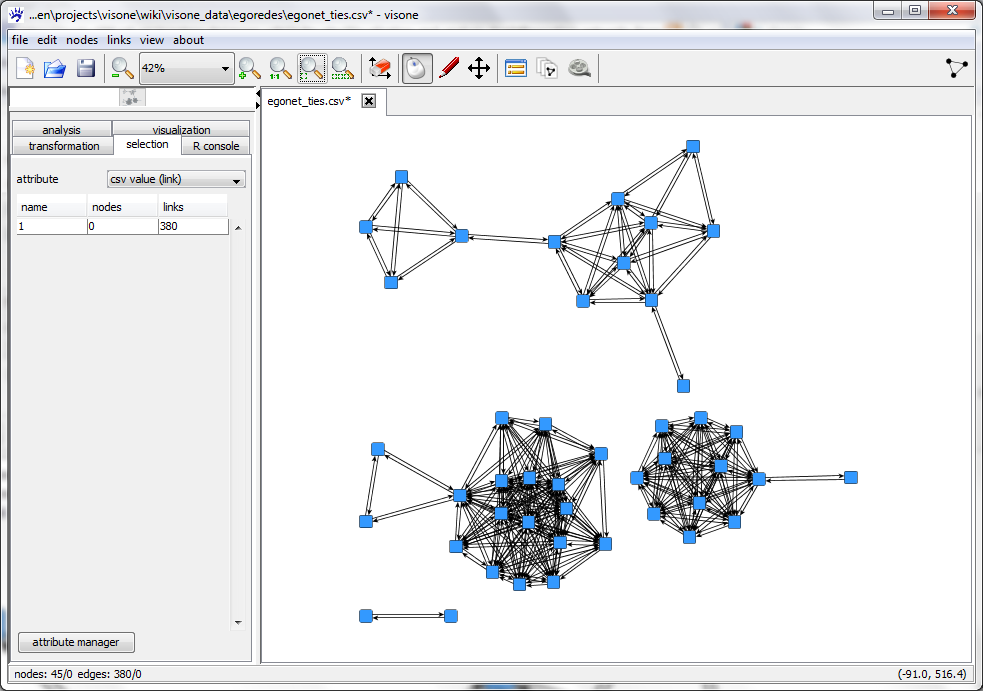
The file view tab shows you (part of) the adjacency matrix encoded in the file to be opened. From this view you can guess, for instance, that different cells in the matrix are delimited by semicolons (;), that row and column labels are present, and some more. For an exhaustive explanation of all options and their meaning see the page on the import options dialog. To continue with this trail, set all options as shown in the format tab above and click on ok. This opens a network looking like this.
The .csv does not contain layout information. The position of the nodes has been determined by the layout algorithm that can be initiated with the quick layout button.
Apart from GraphML and CSV format, visone can also open files in UCINET's .dl format, in Pajek's .net format, and some more.
Merging parallel ties
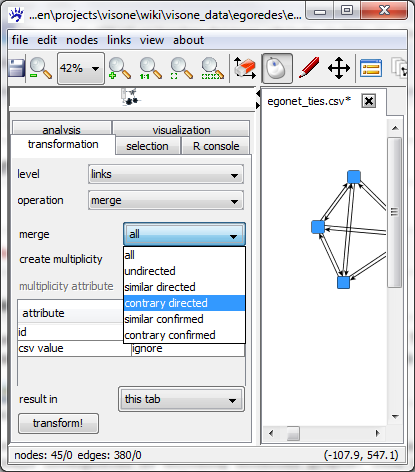
The network above contains for every pair of actors that are connected two anti-parallel ties. This is due to the fact that adjacency matrices are always interpreted as encoding directed graphs. This interpretation is wrong in our example since the tie-generating question was "do actor A and actor B know each other?" which clearly generates an undirected relation. All pairs of anti-parallel directed links can be merged to one undirected link via the transformation tab.
Therefore chose links as the level on which the transformation should be applied, merge as the operation, and chose contrary directed in the drop-down menu right of merge. Clicking on transform! at the bottom of the tab executes the transformation and the network has been transformed into an undirected one with no parallel links.
Since now we have already invested some work in the network, we might save it by clicking on file, save. (The first time we do this we have to assign a name to the network.) Note that the network is saved in GraphML format; indeed only this format guarantees that no information gets lost.
Importing attributes from CSV tables
Currently, the nodes have only one attribute called id. The values of other attributes are provided in the file Egonet_attributes.csv. This file can be merged into our current network via the attribute manager which can be started by clicking on the icon ![]() in visone's toolbar.
in visone's toolbar.
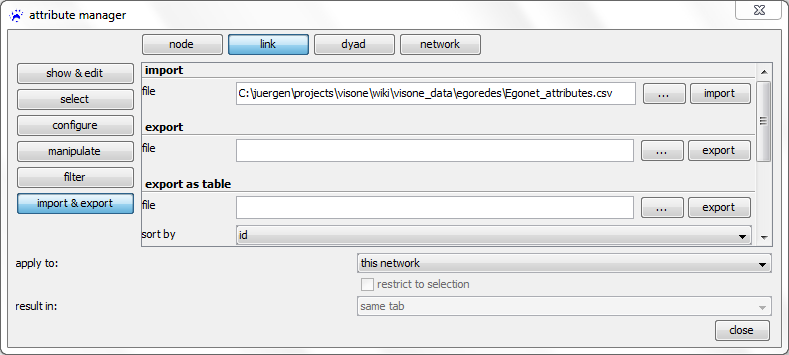
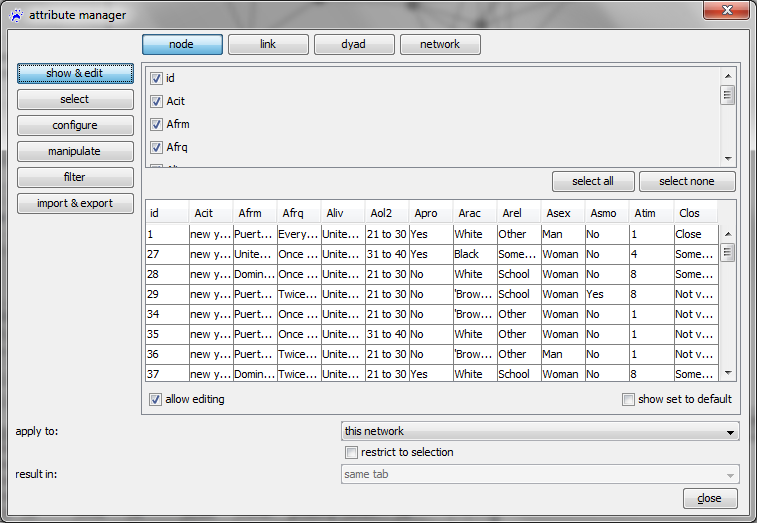
In the attribute manager, choose the nodes radio-button in the top row, import & export on the left, and select the file Egonet_attributes.csv that you have previously downloaded to your computer.
Before clicking on apply it is very important to correctly set the value in the join by drop-down menu. This should point to the name of the attribute that identifies the nodes and tells visone which column in the imported CSV file holds these identifiers. Currently you can only select id but in general there might be several node attributes and the nodes could as well be identified by attributes having a different name than id. Clicking on apply opens an import options dialog; setting the options as shown above - in particular, setting the cell delimiter to semicolon (;) - and clicking on ok imports the attributes. You can see the result by checking the values radio button on the left hand side of the attribute manager. Then, you should see something similar to the image on the right.
Mapping attributes to graphics: categorical attributes
For many networks it is extremly insightful to show the attribute values in the network image. In visone, attributes can be mapped to graphical variables via the visualization tab.
In the visualization tab you can choose between plenty of options; these have to be set from top to bottom. For the category chose mapping since we want to map existing attributes to graphical variables. (The other two options are layout for applying a visualization algorithm and geometry for doing affine transformations, such as rotating or scaling.)
The type of the mapping refers to the type of the graphical variable that is used for encoding attribute values. This choice is restricted by the type of the attribute that is to be mapped. Numerical attributes can be mapped to graphical variables that allow the user to recognize that one actor has a larger value than another one; examples of such variables are size or position whose usage is demonstrated further down in this trail. In this section, however, we want to encode a categorical variable - more specifically, the country of origin of the actor. A good choice for a graphical variable to encode this information is color which we select in the drop-down menu right of type. For property chose node color.
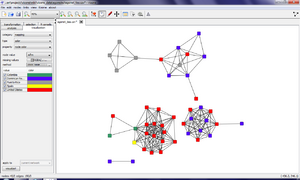
The drop-down menu right of node value should be set to the name of the attribute that is to be encoded; we chose Afrm (meaning country of origin of the actor). Finally, selecting color table for the method option presents you a table with the different values of the Afrm attribute together with a predefined choice of colors that you can change as you wish. Clicking on the visualize! button at the bottom of the visualization tab applies the mapping whose result is shown in the network area of the visone window.
Computing network analytic indicators: centrality
visone offers you a rich choice of methods that compute analytic measures on a given network. In this section, we exemplarily compute a centrality indicator on the set of actors. (Node centralities specify which actors are the important ones with respect to a certain criterion.)
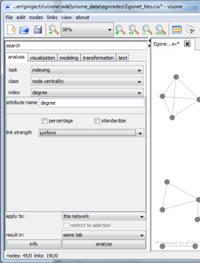
Computing such indicators is done via the analysis tab. To compute the degree of the actors (i.e., the number of ties attached to each actor) set task to indexing. Indexing means that we are assigning values to individual elements (i.e., to nodes or edges) of the network. Set class to node centrality and index to degree. The values specified for link length and link strength can currently only be uniform since we don't have any tie weights. Unchecking the percentage and standardize boxes ensures that the actual degrees are computed. The text field right of result attribute specifies the name of the attribute in which the computed values are to be stored. (visone makes suggestions for this name; at the moment it should say degree but you can change this if you want.)
Having set all these options click on the analyze! button at the bottom of the analysis tab. (You won't see any change in the image of the network; however, at the bottom line of the main window visone tells you what has been computed and in which attribute it has been stored.)
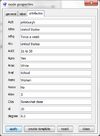
You could browse the values of the newly computed attribute in the attribute manager or in the attributes tab of the node properties dialog (see the last entry telling you that the degree of the selected node equals 9). You could also export the attributes via the attribute manager (which works similar to the import of attributes explained above) and store them in CSV tables.
However, in many situation it is more insightful to visualize the computed centrality values. This is illustrated in the next two sections of this trail.
Mapping attributes to graphics: numerical attributes
Centrality values (such as the node degrees that have been computed above) are numerical attributes. These can be mapped to graphical variables that let the user distinguish between larger and smaller. Examples of such graphics are size, position (x/y-coordinates), or color gradient. For instance, to map the node degree to the node size you can proceed similar as for mapping categorical attributes (illustrated above). In the visualization tab choose category=mapping, type=size, property=node area, and node value=degree; click on visualize!.
Network layout
Network layout algorithms compute the positions of nodes, links, and/or labels to yield nice network images. For convenience, a general-purpose layout algorithm can be initiated by clicking on the quick layout button ![]() in visone's toolbar. All other layout algorithms can be initiated from the visualization tab (select category=layout). The different choices in the drop-down menu right of layout determine which positions are to be modified.
in visone's toolbar. All other layout algorithms can be initiated from the visualization tab (select category=layout). The different choices in the drop-down menu right of layout determine which positions are to be modified.
Selecting elements dependent on attribute values
Finally, we demonstrate how network elements (i.e., nodes and links) that have specific attribute values can be selected. Therefore, download the file Egonet.graphml (see the description on the page Egoredes (data)), save it to your computer, and open it in visone via the menu file, open.
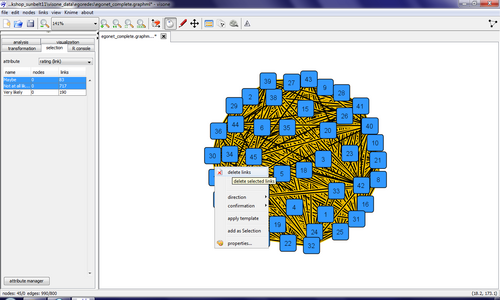
This network turns out to be one big mess containing 45 nodes and 990 links (a simple calculation tells you that the network is complete, i.e., each pair of nodes is connected by a link). The information about which actors actually know each other is encoded in the link attribute rating which can assume the values Not at all likely, Maybe, or Very likely. To delete all links that are not rated as Very likely (say), go to the selection tab and select the attribute rating (link) in the drop-down menu. A list shows the different values of this attribute along with the number of links that are mapped to this value and clicking on one or several rows (keep the Control-Key down to select more than one row) selects these links. The selected links can be deleted either from the link context menu or from the menu links, delete links. Selecting and deleting those links that are rated as Not at all likely, or Maybe and layouting the network via the quick layout button shows you a network that is identical to the one encoded in the adjacency matrix Egonet_ties.csv.
More sophisticated ways to select network elements are illustrated in the next trail on advanced attribute management.